Methylation at the N6 position of adenosine (m6A) is the most abundant internal modification within protein-coding and long noncoding RNAs in eukaryotes, which is a reversible process with important biological functions. YT521-B homology domain family (YTHDF) proteins are the ¢readers¢ of m6A, the binding of which results in the alteration of the translation efficiency and stability of m6A-containing RNAs. However, the mechanism underlay is poorly understood. A research team led by Prof. WU Ligang at National Center for Protein Science·Shanghai, Institute of Biochemistry and Cell Biology (SIBCB), Shanghai Institutes for Biological Sciences (SIBS) of Chinese Academy of Sciences, has uncovered the mechanism of YTHDF2-mediated degradation of m6A-containing RNAs in mammalian cells. This work has been published in Nature Communications. The researchers used a pulse-chase system to monitor the decay of RNAs. They found that both m6A modification and binding of YTHDF2 can destabilized RNA by hastening deadenylation (shortening of the poly(A) tail) as the initial step. Researchers then screened for interactions between YTHDF2 and components of the potential cellular deadenylase complexes. The results drew researchers’ attention to CCR4-NOT, a nine-subunit complex containing two deadenylase subunits (CAF1 and CCR4) and a large scaffold subunit, CNOT1. Dominant-negative assay confirmed that CAF1 and CCR4 are the ribonucleases responsible for the YTHDF2-mediated deadenylation. Further study showed that YTHDF2 recruited the CCR4-NOT complex through a direct interaction between the YTHDF2 N-terminal region and the SH domain of the CNOT1 subunit and that this recruitment was essential for the deadenylation and decay of m6A-containing RNAs. This study provided a further understanding of RNA modifications on regulation of gene expression. This work was jointly conducted Prof. WU Ligang's team and Prof. MA Jinbiao’s lab from School of Life Sciences, Fudan University. This study was supported by the grants from the Ministry of Science and Technology, National Natural Science Foundation of China and China Postdoctoral Science Foundation. 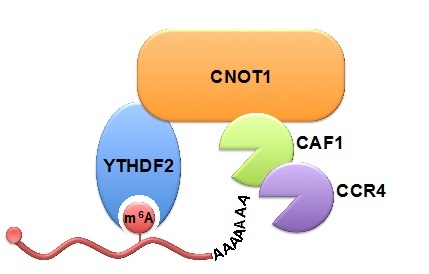
Figure: YTHDF2 binds to the m6A modification on RNA and recruits the CCR4-NOT deadenylase complex through direct interaction with CNOT1. (Image by Prof. WU Ligang's lab) |

